Research in Progress
Current Projects
.png?sfvrsn=9c99b0b4_0)
Surveillance and modeling of infectious diseases in wastewater
Wastewater-based epidemiology (WBE) can provide insights on community spread of endemic and emerging pathogens. During the COVID-19 pandemic, WBE gained widespread popularity and is now a core component of public health surveillance in many areas. Prior research done by our group on COVID-19 surveillance data suggests that wastewater also provides important and unique information for estimating hospital demand.
Our team has partnered with state public health officials and the Colorado Wastewater Surveillance Program to improve the use of WBE to track infectious diseases across Colorado. We conducted a study investigating the use of mobile device and flow data to characterize population fluctuations in wastewater monitoring areas (sewersheds) around the state, and are currently evaluating the performance of different normalization methods for wastewater concentrations of SARS-CoV-2, influenza A, and respiratory syncytial virus (RSV) in predicting hospital admissions at the sewershed level in Colorado. Code is available on GitHub.
.png?sfvrsn=b79ab0b4_0)
Estimating the burden of Long COVID in Colorado
Long COVID has caused substantial morbidity in the United States and in some people, the disease can be life-altering; however, the burden of disease has been challenging to estimate. This is due in part to the fact that the disease is new and only recently was a clear definition of Long COVID established, leading to underdiagnosis. Our team has partnered with the Colorado Department of Public Health & Environment and the Lieutenant Governor's Office to estimate the burden and costs of Long COVID in Colorado. We are using health insurance claims data from the Colorado All Payer Claims Database (APCD) to identify healthcare utilization patterns among those diagnosed with Long COVID. We are also developing predictive algorithms using machine learning to estimate the prevalence of undiagnosed Long COVID in Colorado using these claims data, and evaluating the distribution of disease across the state.
.png?sfvrsn=199eb0b4_0)
Characterizing extreme weather drivers of H5N1 (avian influenza)
Highly Pathogenic Avian Influenza (HPAI), particularly H5N1, has emerged as a pathogen of global concern. Since its initial emergence in 1959, H5N1 has caused widespread mortality in wild birds, large-scale outbreaks and culling events in the poultry industry, and recurrent and increasing spillover events that pose significant pandemic threats. HPAI H5N1 was first detected in wild bird populations in North America in November 2021, which has already resulted in over 15,000 cases from more than 160 North American wild bird species, and evidence of spillover to over 400 wild mammals from over 20 distinct species, including 71 reported human infections and one death in the US. One potential yet under-explored reason for rising spillover and outbreaks events is that shifting patterns in temperature, precipitation, and the frequency and intensity of Extreme Weather Events (EWEs) are altering ecological interactions at the wildlife-agriculture-human interface, influencing where and when spillover occurs.
We received a pilot award from the BUSPH-HSPH-CAFE, funded by NIH U24ES035309, to develop predictive models aimed at understanding the risk of H5N1 outbreaks in commercial poultry, backyard flocks, and wild birds across the United States using a combination of bird migration, weather and storm, and EWE data. This work was recently posted as a preprint.
.png?sfvrsn=6b9fb0b4_0)
Using climate and ecological information to improve tools to estimate West Nile Virus risk
West Nile virus transmission is shaped by complex interactions among temperature, humidity, and precipitation patterns, mosquito vectors, avian host populations, and local environmental conditions. Recent severe outbreaks in the US, especially in Colorado, underscore the importance of improving our understanding of the ecological pathways linking environmental factors to human West Nile virus risk. Our work integrates weather, ecological land cover, and surveillance data across US regions to identify periods and locations of elevated outbreak risk and to improve forecasting tools that can better support public health preparedness.

Using epidemiology, ecology, and genomics to advance elimination of schistosomiasis in China
Several areas around the world, including China, are aiming to eliminate schistosomiasis, but attaining this goal has been challenging, as despite aggressive control measures, the disease can reemerge and persist in areas for reasons not fully understood. Schistosomiasis is a waterborne pathogen that causes liver fibrosis and anemia and can impair child growth and development. Since 2006, we have worked with collaborators at the Sichuan Center for Disease Control and Prevention to identify sources of infection in schistosomiasis "hotspots" and to improve surveillance tools for areas approaching elimination. In our research, we found that the conditions driving infection risk shift over time and become increasingly localized, and demonstrated that infections tend to be concentrated in a limited number of individuals. We found that the agricultural practice of using human and animal waste as an agricultural fertilizer (often called night soil) may facilitate transmission in human populations and bovine populations. We developed an efficient method for sequencing large numbers of loci from field-collected S. japonicum miracidia and have shown how genomic tools can be used to improve schistosomiasis tools in theory and in practice. We have employed remote sensing and geospatial analytical tools to evaluate best-practice schistosomiasis surveillance efforts within low-transmission environments. We evaluated the role of travel in schistosomiasis exposure and infection risk (in our study villages where some people regularly travel to cities, we found more travel was associated with reduced water contact and infection risk). Current efforts are underway to understand the roles of parasite import and export along social and hydrological gradients and evaluate bovines as sources of infection in residual transmission hotspots. This research has been funded by the National Institute of Allergy and Infectious Diseases.

Impacts of climate change on waterborne disease
There is strong evidence that climate impacts the distribution of waterborne diseases. The causal pathways are complex, and health impacts of climate change depend not only on meteorological exposures, but also other underlying vulnerabilities. We are interested in improving our understanding of these causal pathways so that we can identify locations and time windows of elevated risk and, ultimately, design interventions and early warning systems to reduce the burden of waterborne diseases in a changing climate. Our early work in this area includes a systematic review, a meta-analysis, and a proposed framework to characterize the relationships between climate drivers, including temperature, heavy rainfall, flooding, and drought, and their relationship with infectious diarrhea. More recently, our work has included a study estimating the impact of weather on the use of safe and unsafe drinking water in countries in Asia and Africa, and a study on the impact of precipitation and temperature on giardia and cryptosporidiosis in Colorado.
Past Projects
.jpg?sfvrsn=f43bc6b9_0)
Supporting the COVID-19 pandemic response
At the onset of the COVID-19 pandemic in March 2020, our group became a core part of the Colorado COVID-19 Modeling Group. This team used modeling, data analysis, and rapid evidence synthesis to inform state and local officials in Colorado about the current state and possible future trajectories of the pandemic. In 2022, with funding from the Council of State and Territorial Epidemiologists (CSTE), our team expanded its focus to developing modeling tools and data visualizations to meet the needs of public health leaders in the Rocky Mountain West (RMW). We launched and maintained a website from the end of the COVID-19 public health emergency (PHE) through December 2023, providing interactive COVID-19 data visualizations and summaries of emerging topics for COVID-19 planning and response in the RMW. This website was developed following interviews with public health officials across the RMW region to assess their needs during a transitional period in the pandemic response.
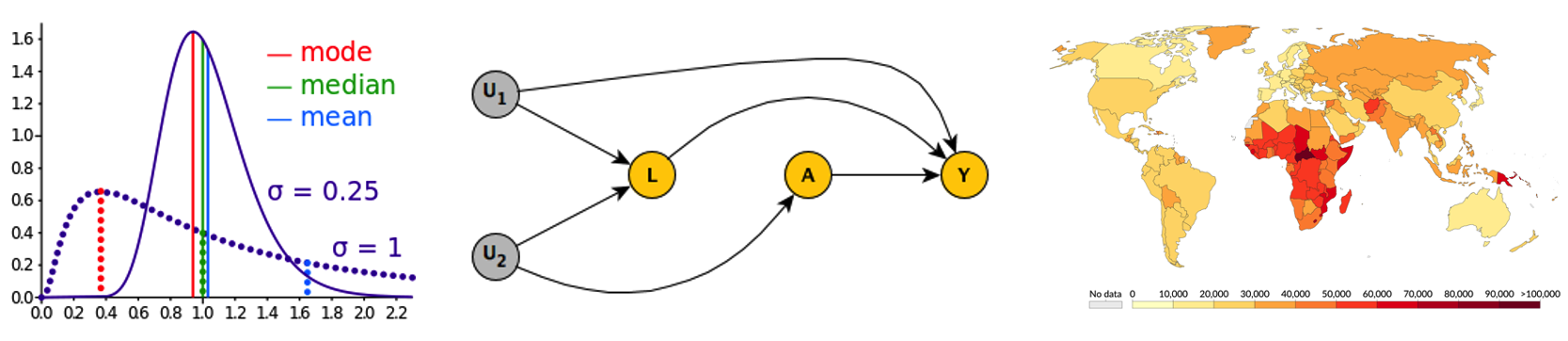
Training for communicable disease epidemiologists
The Carlton Research Group developed training seminars for communicable disease epidemiologists at state and local health departments around Colorado. Topics included core concepts in epidemiology, biostatistics, the health impacts of climate change, and the use of mathematical modeling to inform pandemic policy, with worked examples using communicable diseases. Our aim was to create engaging educational experiences through trainings that included foundational concepts, real-world examples, and connection to additional resources for further learning. For more information on trainings given by our research group, please contact Emma Gunn at [email protected].


